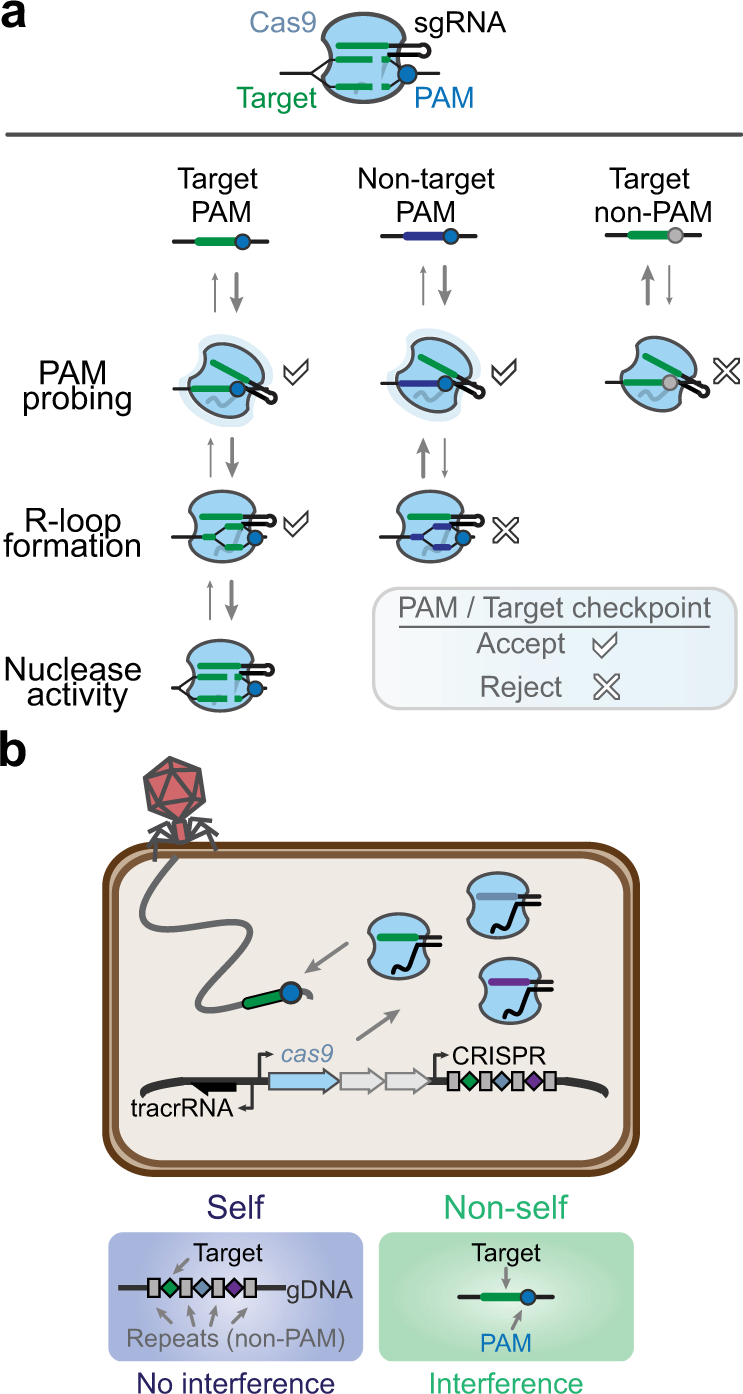
Mechanistic details of CRISPR-associated transposon recruitment and integration revealed by cryo-EM | PNAS
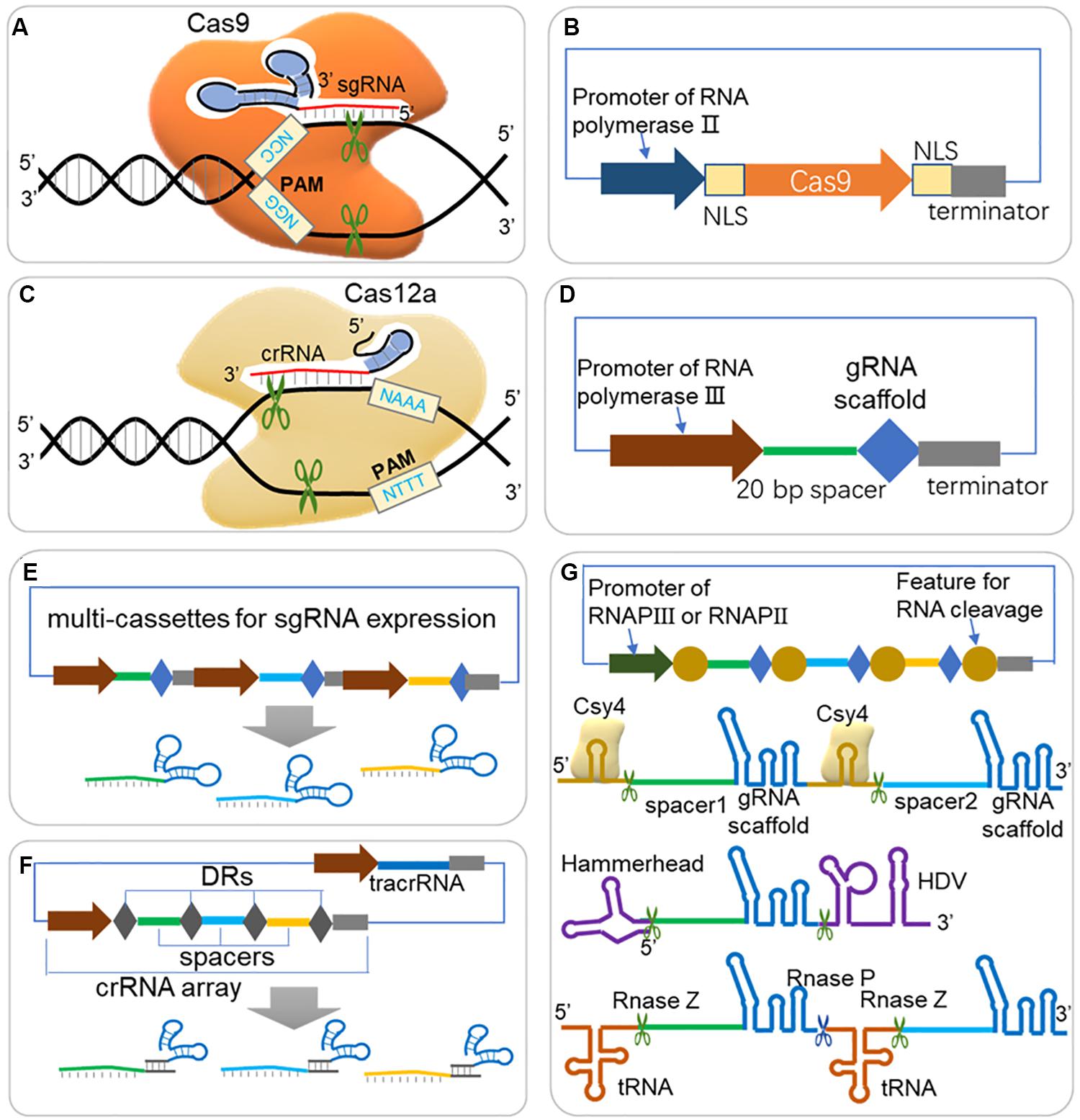
![PDF] Harnessing “A Billion Years of Experimentation”: The Ongoing Exploration and Exploitation of CRISPR–Cas Immune Systems | Semantic Scholar PDF] Harnessing “A Billion Years of Experimentation”: The Ongoing Exploration and Exploitation of CRISPR–Cas Immune Systems | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/b22a039b9f9886dbfd310bc92de89a097b8e2262/9-Figure3-1.png)
PDF] Harnessing “A Billion Years of Experimentation”: The Ongoing Exploration and Exploitation of CRISPR–Cas Immune Systems | Semantic Scholar

Efficient Bacterial Genome Engineering throughout the Central Dogma Using the Dual-Selection Marker tetAOPT | ACS Synthetic Biology

IIC Partners on Twitter: "Harry Klompe of Danone speaks on Poland's talent market #iicpartners #executivesearch #warsaw https://t.co/1Dd0iXSSNK" / Twitter
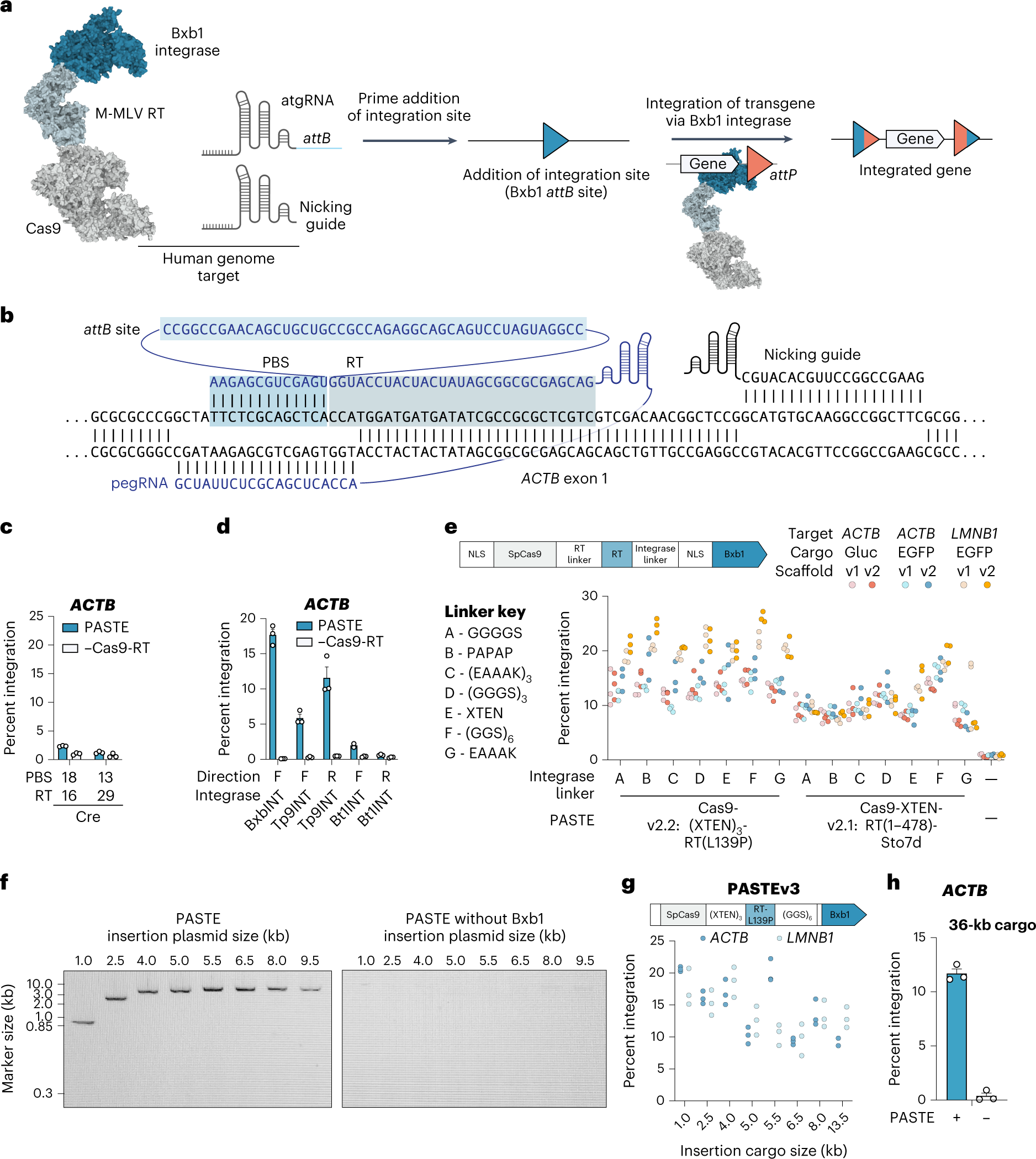
Drag-and-drop genome insertion of large sequences without double-strand DNA cleavage using CRISPR-directed integrases | Nature Biotechnology

How to promote multifunctionality of vegetated strips in arable farming: A qualitative approach for Germany - Schütz - 2022 - Ecosphere - Wiley Online Library